Understandings:
Applications and skills:
Guidance:
- Ribosomes
Once mRNA is produced from the DNA template, the process of actually producing the protein at the ribosomes can begin.
This process is referred to as translation because it changes the language of DNA to the language of protein. The centre of this process is the ribosome. Therefore, you need to understand the structure of this organelle.
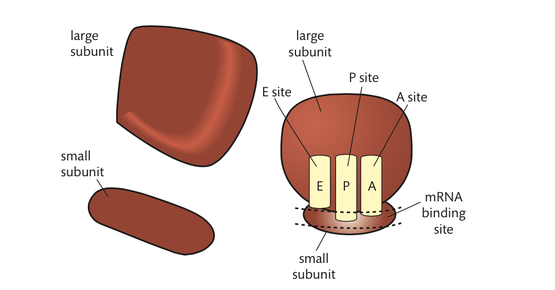
Ribosomes can be seen with an electron microscope. Each ribosome consists of a large subunit and a small subunit. The subunits are composed of ribosomal RNA (rRNA) molecules and many distinct proteins. rRNA proteins are generally small and are associated with the core of the RNA subunits. Roughly two-thirds of their mass is rRNA. The molecules of the ribosomes are constructed in the nucleolus of eukaryotic cells and exit the nucleus through the membrane pores. Prokaryotic ribosomes are smaller than eukaryotic ribosomes. There is also a difference in molecular makeup.
The decoding of a strand of mRNA to produce a polypeptide occurs in the space between the two subunits. In this area, there are binding sites for mRNA and three sites for the binding of tRNA
- Initiation of translation involves assembly of the components that carry out the process.
- Synthesis of the polypeptide involves a repeated cycle of events.
- Disassembly of the components follows termination of translation.
- Free ribosomes synthesize proteins for use primarily within the cell.
- Bound ribosomes synthesize proteins primarily for secretion or for use in lysosomes.
- Translation can occur immediately after transcription in prokaryotes due to the absence of a nuclear membrane.
- The sequence and number of amino acids in the polypeptide is the primary structure.
- The secondary structure is the formation of alpha helices and beta pleated sheets stabilized by hydrogen bonding.
- The tertiary structure is the further folding of the polypeptide stabilized by interactions between R-groups.
- The quaternary structure exists in proteins with more than one polypeptide chain.
Applications and skills:
- Application: tRNA-activating enzymes illustrate enzyme–substrate specificity and the role of phosphorylation.
- Skill: Identification of polysomes in electron micrographs of prokaryotes and eukaryotes.
- Skill: The use of molecular visualization software to analyse the structure of eukaryotic ribosomes and a tRNA molecule.
Guidance:
- Names of the tRNA binding sites are expected, as well as their roles.
- Examples of stop and start codons are not required.
- Polar and non-polar amino acids are relevant to the bonds formed between R-groups.
- Quaternary structure may involve the binding of a prosthetic group to form a conjugated protein.
- Ribosomes
Once mRNA is produced from the DNA template, the process of actually producing the protein at the ribosomes can begin.
This process is referred to as translation because it changes the language of DNA to the language of protein. The centre of this process is the ribosome. Therefore, you need to understand the structure of this organelle.
Ribosomes can be seen with an electron microscope. Each ribosome consists of a large subunit and a small subunit. The subunits are composed of ribosomal RNA (rRNA) molecules and many distinct proteins. rRNA proteins are generally small and are associated with the core of the RNA subunits. Roughly two-thirds of their mass is rRNA. The molecules of the ribosomes are constructed in the nucleolus of eukaryotic cells and exit the nucleus through the membrane pores. Prokaryotic ribosomes are smaller than eukaryotic ribosomes. There is also a difference in molecular makeup.
The decoding of a strand of mRNA to produce a polypeptide occurs in the space between the two subunits. In this area, there are binding sites for mRNA and three sites for the binding of tRNA
The triplet bases of the mRNA codon pair with the complementary bases of the triplet anticodon of the tRNA.
Polypeptide chains are assembled in the cavity between the two subunits. This area is generally free of proteins, so binding of mRNA and tRNA is carried out by the rRNA. tRNA moves sequentially through the three binding sites: from the A site, to the P site, and finally to the E site. The growing polypeptide chain exits the ribosome through a tunnel in the large subunit core.
- Translation: RNA → protein
The translation process involves several phases:
• initiation • translocation • elongation • termination.
Before we discuss these phases, it is important to consider the codons. You will recall that codons carry the genetic code from DNA to the ribosomes via mRNA. There are 64 possible codons. Three codons have no complementary tRNA anticodon: these are the stop codons. There is a start codon (AUG) that signals the beginning of a polypeptide chain. This codon also encodes the amino acid methionine.
- The initiation phase
The start codon (AUG) is on the 5ʹ end of all mRNAs. Each codon, other than the three stop codons, attaches to a particular tRNA. The tRNA has a 5ʹ and a 3ʹ end, like all other nucleic acid strands. The 3ʹ end of tRNA is free and has the base sequence CCA. This is the site of amino acid attachment. Because there are complementary bases in the single-stranded tRNA, hydrogen bonds form at four areas. This causes the tRNA to fold and take on a three-dimensional structure. If the molecule is flattened, it has a two-dimensional appearance resembling a three-leaf clover. One of the loops of the clover leaf contains an exposed anticodon. This anticodon is unique to each type of tRNA. It is this anticodon that pairs with a specific codon of mRNA.
The first step in initiation of translation is when an activated amino acid, methionine attached to a tRNA with the anticodon UAC, combines with an mRNA strand and a small ribosomal subunit.
The small subunit moves down the mRNA until it contacts the start codon (AUG). This contact starts the translation process. Hydrogen bonds form between the initiator tRNA and the start codon. Next, the large ribosomal subunit combines with these parts to form the translation initiation complex. Joining the initiation complex are proteins called initiation factors that require energy from guanosine triphosphate (GTP) for attachment. GTP is an energy-rich compound very similar to ATP.
- The elongation phase
Once initiation is complete, elongation occurs. This phase involves tRNAs bringing amino acids to the mRNA–ribosomal complex in the order specified by the codons of the mRNA. Proteins called elongation factors assist in binding the tRNAs to the exposed mRNA codons at the A site. The initiator tRNA moves to the P site. The ribosomes catalyse the formation of peptide bonds between adjacent amino acids that are brought to the polypeptide assembling area.
- The translocation phase
The translocation phase actually happens during the elongation phase. Translocation involves the movement of the tRNAs from one site of the mRNA to another. First, a tRNA binds with the A site. Its amino acid is then added to the growing polypeptide chain by a peptide bond. This causes the polypeptide chain to be attached to the tRNA at the A site. The tRNA then moves to the P site. It transfers its polypeptide chain to the new tRNA, which moves into the now exposed A site. The now empty tRNA is transferred to the E site, where it is released. This process occurs in the 5ʹ to 3ʹ direction. Therefore, the ribosomal complex is moving along the mRNA towards the 3ʹ end. Remember, the start codon was near the 5ʹ end of the mRNA.
- The termination phase
The termination phase begins when one of the three stop codons appears at the open A site. A protein called a release factor then fills the A site. The release factor does not carry an amino acid. It catalyses hydrolysis of the bond linking the tRNA in the P site with the polypeptide chain. This frees the polypeptide, releasing it from the ribosome. The ribosome then separates from the mRNA and splits into its two subunits.
The termination phase completes the process of translation. At this point, a disassembly process occurs in which the mRNA detaches from the ribosome, all tRNAs detach from the mRNA–ribosomal complex, and the protein is released from the ribosome.
- Polar and non-polar amino acids
Amino acids are often grouped based on the properties of their side chains (R-groups). Amino acids with non-polar side chains are hydrophobic. Non-polar amino acids
are found in the regions of proteins that are linked to the hydrophobic area of the cell membrane.
Polar amino acids have hydrophilic properties, and they are found in regions of proteins that are exposed to water. Membrane proteins include polar amino acids towards the interior and exterior of the membrane. These amino acids create hydrophilic channels in proteins through which polar substances can move.
Polar and non-polar amino acids are important in determining the specificity of an enzyme. Each enzyme has a region called the active site. Only specific substrates can combine with particular active sites. Combination is possible when fitting occurs. Fitting involves the general shapes and polar properties of the substrate and of the amino acids exposed at the active site.
Polypeptide chains are assembled in the cavity between the two subunits. This area is generally free of proteins, so binding of mRNA and tRNA is carried out by the rRNA. tRNA moves sequentially through the three binding sites: from the A site, to the P site, and finally to the E site. The growing polypeptide chain exits the ribosome through a tunnel in the large subunit core.
- Translation: RNA → protein
The translation process involves several phases:
• initiation • translocation • elongation • termination.
Before we discuss these phases, it is important to consider the codons. You will recall that codons carry the genetic code from DNA to the ribosomes via mRNA. There are 64 possible codons. Three codons have no complementary tRNA anticodon: these are the stop codons. There is a start codon (AUG) that signals the beginning of a polypeptide chain. This codon also encodes the amino acid methionine.
- The initiation phase
The start codon (AUG) is on the 5ʹ end of all mRNAs. Each codon, other than the three stop codons, attaches to a particular tRNA. The tRNA has a 5ʹ and a 3ʹ end, like all other nucleic acid strands. The 3ʹ end of tRNA is free and has the base sequence CCA. This is the site of amino acid attachment. Because there are complementary bases in the single-stranded tRNA, hydrogen bonds form at four areas. This causes the tRNA to fold and take on a three-dimensional structure. If the molecule is flattened, it has a two-dimensional appearance resembling a three-leaf clover. One of the loops of the clover leaf contains an exposed anticodon. This anticodon is unique to each type of tRNA. It is this anticodon that pairs with a specific codon of mRNA.
The first step in initiation of translation is when an activated amino acid, methionine attached to a tRNA with the anticodon UAC, combines with an mRNA strand and a small ribosomal subunit.
The small subunit moves down the mRNA until it contacts the start codon (AUG). This contact starts the translation process. Hydrogen bonds form between the initiator tRNA and the start codon. Next, the large ribosomal subunit combines with these parts to form the translation initiation complex. Joining the initiation complex are proteins called initiation factors that require energy from guanosine triphosphate (GTP) for attachment. GTP is an energy-rich compound very similar to ATP.
- The elongation phase
Once initiation is complete, elongation occurs. This phase involves tRNAs bringing amino acids to the mRNA–ribosomal complex in the order specified by the codons of the mRNA. Proteins called elongation factors assist in binding the tRNAs to the exposed mRNA codons at the A site. The initiator tRNA moves to the P site. The ribosomes catalyse the formation of peptide bonds between adjacent amino acids that are brought to the polypeptide assembling area.
- The translocation phase
The translocation phase actually happens during the elongation phase. Translocation involves the movement of the tRNAs from one site of the mRNA to another. First, a tRNA binds with the A site. Its amino acid is then added to the growing polypeptide chain by a peptide bond. This causes the polypeptide chain to be attached to the tRNA at the A site. The tRNA then moves to the P site. It transfers its polypeptide chain to the new tRNA, which moves into the now exposed A site. The now empty tRNA is transferred to the E site, where it is released. This process occurs in the 5ʹ to 3ʹ direction. Therefore, the ribosomal complex is moving along the mRNA towards the 3ʹ end. Remember, the start codon was near the 5ʹ end of the mRNA.
- The termination phase
The termination phase begins when one of the three stop codons appears at the open A site. A protein called a release factor then fills the A site. The release factor does not carry an amino acid. It catalyses hydrolysis of the bond linking the tRNA in the P site with the polypeptide chain. This frees the polypeptide, releasing it from the ribosome. The ribosome then separates from the mRNA and splits into its two subunits.
The termination phase completes the process of translation. At this point, a disassembly process occurs in which the mRNA detaches from the ribosome, all tRNAs detach from the mRNA–ribosomal complex, and the protein is released from the ribosome.
- Polar and non-polar amino acids
Amino acids are often grouped based on the properties of their side chains (R-groups). Amino acids with non-polar side chains are hydrophobic. Non-polar amino acids
are found in the regions of proteins that are linked to the hydrophobic area of the cell membrane.
Polar amino acids have hydrophilic properties, and they are found in regions of proteins that are exposed to water. Membrane proteins include polar amino acids towards the interior and exterior of the membrane. These amino acids create hydrophilic channels in proteins through which polar substances can move.
Polar and non-polar amino acids are important in determining the specificity of an enzyme. Each enzyme has a region called the active site. Only specific substrates can combine with particular active sites. Combination is possible when fitting occurs. Fitting involves the general shapes and polar properties of the substrate and of the amino acids exposed at the active site.